Portal:AOP
From WikiPathways
The Adverse Outcome Pathway (AOP) Portal on WikiPathways
Welcome to the Adverse Outcome Pathway Portal!
This Adverse Outcome Pathway (AOP) portal for WikiPathways is created to highlight the molecular basis of AOPs or events in AOPS. In general, AOPs start with a Molecular Initiating Event (MIE) caused by a stressor, followed by Key Events (KEs), that lead to an Adverse Outcome (AO). These AOPs are intended specifically for regulatory decision making and are typically stored in the AOP Knowledge Base (AOPKB). Because AOPs are simplified explanations of biological effects after the effect of a stressor they are not useful to describe and understand the molecular basis of the AOPs and not suited to do data analysis. Such analysis is needed especially for in silico risk analysis intending to lower animal use in toxicology studies. This portal was created to present this previously missing molecular level of the AOPs and get more into detail on the biological processes involved in them. The development of these molecular AOPS is relevant for the European research projects on toxicology EU-ToxRisk and OpenRiskNet that also funded part of the work. This portal is also featured in the latest NAR Database Issue on WikiPathways.
News
2021-07-09 - Formalised the molecular AOP development protocol and added brief description in the Mission section
2021-01-06 - Started development of COVID-19 AOP networks linked to ongoing efforts in the CIAO project.
2020-08-07 - Added new format of molecular AOPs shaped as meta-pathways, including DataNodes for AOP-Wiki Key Events, including linkout
2018-12-21 - new paper: Introducing WikiPathways as a Data-Source to Support Adverse Outcome Pathways for Regulatory Risk Assessment of Chemicals and Nanomaterials
2018-04-04 - Changed Featured Pathways page, now shows images of all pathways included in the portal
2017-08-14 - Proposed AOP list added to the Mission section
2017-05-19 - The first AOP based on descriptive text from aopwiki.org on liver fibrosis is added to the portal.
2017-03-29 - The first AOPs that are added into this portal on pulmonary fibrosis, are results of the eNanoMapper project.
2017-03-29 - This portal was created
Mission
Purpose
The purpose of this portal is to create a collection of AOPs on the molecular level and facilitating the use of high-throughput transcriptomic data in the AOP framework. The subjects of the first AOPs are linked to the use cases of the EU-ToxRisk program, based on [aopwiki.org AOP-Wiki] entries, and the CIAO project on COVID-19 AOPs.
List of molecular AOPs present in this portal (some are more defined than others):
- Mitochondrial Complex I inhibition leads to liver injury
- Mitochondrial Complex I inhibition leads to parkinsonian motor deficits
- Mitochondrial Complex I inhibition leads to Fanconi syndrome
- Protein alkylation leads to liver fibrosis
- Interaction with lung resident cell membrane components leads to lung fibrosis
- Multiple MIEs lead to liver steatosis
- ACE2 inhibition leads to increased mortality
Basic strategies and principles for general AOPs are described in this paper:
Villeneuve et al. (2014). Adverse Outcome Pathway (AOP) Development I: Strategies and Principles. Toxicological Sciences PubMed
The list of Featured Pathways is not static and can be updated at any time. If you know of a pathway that should be added, please contact Marvin Martens.
Creating molecular AOPs
Molecular AOPs are developed as so-called meta-pathways, in which the overall layout of the AOP is preserved as in literature or AOP-Wiki in terms of Key Events. These exist as new "Event" type datanodes containing AOP-Wiki identifiers, and are connected with basic directed interactions. Next, molecular pathways of processes described in Key Events (existing in WikiPathways database or newly developed) are connected with the Event nodes with basic interactions. This structure allows use of the AOP's modular nature to generate AOP networks and extend with molecular entities such as genes, proteins and metabolites using Cytoscape and the apps of WikiPathways and CyTargetLinker.
Featured Pathway
|
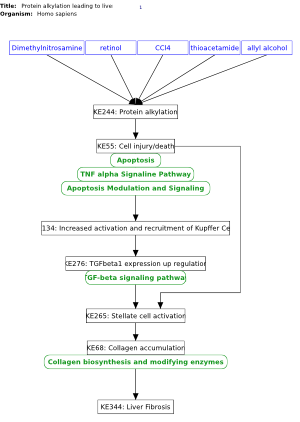
Protein alkylation leading to liver fibrosis (Homo sapiens) |
| View all Featured Pathways for this Portal |
Support
This project has received funding from the European Union’s Horizon 2020 (EU 2020) research and innovation program under grant agreement no. 681002 (EU-ToxRisk), EINFRA-22-2016 program under grant agreement no. 731075 (OpenRiskNet), and NWA grant no. 1292.19.272 (VHP4Safety)