Help:Guidelines
From WikiPathways
| Line 8: | Line 8: | ||
* Pathway names should be in English. | * Pathway names should be in English. | ||
* Pathways should have a descriptive name, like “Cholesterol Biosynthesis”. | * Pathways should have a descriptive name, like “Cholesterol Biosynthesis”. | ||
| - | * Do not include your name or username in the pathway name. | + | * Do not include your name or username in the pathway name. Relevant species name or species codes are also not recommended in pathway names. |
* Try to pick a name that is unique at WikiPathways, to avoid having multiple pathways with the same name. | * Try to pick a name that is unique at WikiPathways, to avoid having multiple pathways with the same name. | ||
* If your pathway is specific to a particular state or tissue, this may be included in the name, for example as a prefix or suffix. | * If your pathway is specific to a particular state or tissue, this may be included in the name, for example as a prefix or suffix. | ||
| Line 55: | Line 55: | ||
* On the other hand, common entities that are involved in many reactions may be drawn as multiple entities for clarity, so you don't get a large messy hairball. Common metabolites include H2O, H+, ATP, NADH. | * On the other hand, common entities that are involved in many reactions may be drawn as multiple entities for clarity, so you don't get a large messy hairball. Common metabolites include H2O, H+, ATP, NADH. | ||
* Certain common metabolites that are unimportant for understanding of the pathway as a whole, such as H2O and H+ may even be left out altogether. | * Certain common metabolites that are unimportant for understanding of the pathway as a whole, such as H2O and H+ may even be left out altogether. | ||
| + | ==== Representing enzymatic reactions ==== | ||
| + | In order to properly store enzymatic reactions in the WikiPathways data model, place a line anchor on the line connecting the two reactants, then use a second connector from the enzyme to the anchor to denote the enzymatic interaction. To add an anchor to a line, right-click on the line and select 'Add Anchor'. | ||
| + | Once the reaction has been drawn as described, the enzyme entity can be placed on top of the connector between reactants, to resemble a typical enzymatic reaction drawing. While the connector to the anchor point is still there, it is effectively "hidden" under the enzyme. | ||
| + | {|class="prettytable" | ||
| + | |- | ||
| + | !Enzymatic reaction | ||
| + | !Enzymatic reaction, connector is hidden | ||
| + | |- | ||
| + | |[[Image:Enzyme.png]] | ||
| + | |[[Image:Enzyme-hidden.png]] | ||
| + | |} | ||
==== Stoichiometry ==== | ==== Stoichiometry ==== | ||
The stoichiometry of reactions is not usually modeled in wikipathways. If you wish you can add this information as a comment attached to a reaction or you can use the label for a metabolite to indicate stoichiometry, as suggested below. | The stoichiometry of reactions is not usually modeled in wikipathways. If you wish you can add this information as a comment attached to a reaction or you can use the label for a metabolite to indicate stoichiometry, as suggested below. | ||
Revision as of 19:32, 5 November 2009
When creating or editing pathway information at WikiPathways, please keep the following guidelines in mind. These guidelines aim to promote consistency, accuracy and stability for the pathway content. By following these guidelines, you will help to maintain the highest possible quality of pathway information and to promote the success of community-based curation.
Contents[hide] |
General guidelines
Pathway naming
- Pathway names should be in English.
- Pathways should have a descriptive name, like “Cholesterol Biosynthesis”.
- Do not include your name or username in the pathway name. Relevant species name or species codes are also not recommended in pathway names.
- Try to pick a name that is unique at WikiPathways, to avoid having multiple pathways with the same name.
- If your pathway is specific to a particular state or tissue, this may be included in the name, for example as a prefix or suffix.
- If the pathway is based on content from an external source, the name of the source may be included in the name, for example as a prefix or suffix.
Author information on pathways
When creating a new pathway, your name as designated in your profile will automatically be added to the Pathway Page. To ensure that your information is linked to the pathway even if it is downloaded and used in external applications, you should also add your name and contact information to the Information Area of the pathway. This is done in the editor by selecting the Information Area and editing the relevant information in the Properties panel.
Pathway annotation
Properly annotating pathways is essential to maintaining a high quality of curation and content at WikiPathways. Annotation includes assigning the pathway to one or more categories, defining each node with a database ID and adding a description for each pathway.
References to literature
Citing peer-reviewed publications is a quality standard in the scientific community and WikiPathways is no exception. We strongly encourage including relevant literature references both at the pathway level (Bibliography) and for individual interactions in pathways.
References to other sources
For all evidence from sources other than the scientific literature, please cite the source in the Bibliography section.
Making major edits
When making major edits (deletions in particular) to an existing pathway not created by you, first suggest the changes on the pathway discussion page, as a courtesy to the original author. Starting a discussion about the proposed changes is a productive way of community editing.
Content forking
If you plan to add tissue- or state-specific information to a pathway describing a general process, please create a new pathway and copy the relevant content from the original pathway, rather than making edits directly to the original pathway. When naming the new related pathway, it is appropriate to use the name of the original pathway, with a suffix or addition explaining the edits. This will ensure that pathway names are not duplicated and will help in organizing related pathway content.
Pathway boundaries
Multiple pathways may overlap or correspond to the same topic, but each should serve a unique goal in what it represents. First browse the current contents at WikiPathways to avoid duplicate work. Browsing related pathways can also give you an idea of appropriate pathway boundaries.
Other curator guidelines
Guidelines developed by other resources can also be consulted. While much of the material may be specific to their curation tools, there are general rules that apply to any pathway curation effort... where sloppy biology must be systematically modeled and ultimately displayed on a page.
- Peter Karp and the BioCyc group provide an excellent and thorough specification to help guide all aspects of the curation of metabolic pathways: BioCyc guidelines.
- Reactome hosts a wiki where they've posted the Reactome Curator Guide
- Kitano H, Funahashi A, Matsuoka Y, Oda K: Using process diagrams for the graphical representation of biological networks. Nat Biotechnol 2005, 23(8):961-966. pubmed
- Pirson I, Fortemaison N, Jacobs C, Dremier S, Dumont JE, Maenhaut C: The visual display of regulatory information and networks. Trends Cell Biol 2000, 10(10):404-408. pubmed
- Kohn KW, Aladjem MI, Weinstein JN, Pommier Y: Molecular interaction maps of bioregulatory networks: a general rubric for systems biology. Mol Biol Cell 2006, 17(1):1-13. pubmed
Graphical style guidelines
It is up to the individual author of a pathway to create a style that best displays the relevant information. WikiPathways does not enforce a particular graphical style for pathways. Here we offer a few suggestions on specific aspects of graphical style that best utilize the tools and data model.
Representing complexes
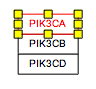
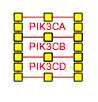
Representing molecular complexes can be done in a variety of styles. We recommend using a stacked representation whenever possible, which can easily be accomplished with the “Stack genes” feature in the editor Toolbar. The example below displays a stacked representation of the complex of subunits A, B, G and D1 of the human DNA polymerase.
You will also likely want to "group" the stacked genes. Once selected, genes can be grouped by pressing CTRL-G, or choosing 'group' in the right-click menu
| Before | After |
|---|---|
| 
|
Representing reactions
- When two reaction steps follow each other in a pathway (i.e. the product of reaction A is the substrate for reaction B), it is best to use a single element for this entity.
- On the other hand, common entities that are involved in many reactions may be drawn as multiple entities for clarity, so you don't get a large messy hairball. Common metabolites include H2O, H+, ATP, NADH.
- Certain common metabolites that are unimportant for understanding of the pathway as a whole, such as H2O and H+ may even be left out altogether.
Representing enzymatic reactions
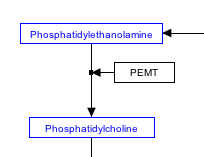
In order to properly store enzymatic reactions in the WikiPathways data model, place a line anchor on the line connecting the two reactants, then use a second connector from the enzyme to the anchor to denote the enzymatic interaction. To add an anchor to a line, right-click on the line and select 'Add Anchor'. Once the reaction has been drawn as described, the enzyme entity can be placed on top of the connector between reactants, to resemble a typical enzymatic reaction drawing. While the connector to the anchor point is still there, it is effectively "hidden" under the enzyme.
| Enzymatic reaction | Enzymatic reaction, connector is hidden |
|---|---|
| 
|
Stoichiometry
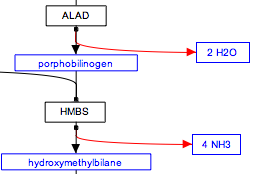
The stoichiometry of reactions is not usually modeled in wikipathways. If you wish you can add this information as a comment attached to a reaction or you can use the label for a metabolite to indicate stoichiometry, as suggested below.
| Stoichiometry in metabolite labels |
|---|
|
Representing multiple similar reactions
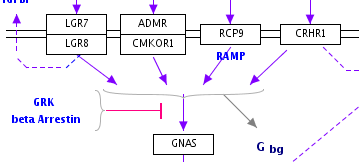
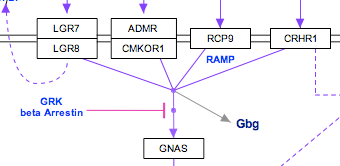
Representing multiple similar reactions so that they are properly stored in the underlying data model can sometimes present a challenge. Below is an example of using a connector hub (a line anchor) to display this. Right-click on any line to add a line anchor, then connect multiple lines for different reactions to this hub.
| Wrong | Right! |
|---|---|
| 
|
Using labels
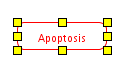
To enter text for a label, double-click on the label (or choose 'Properties' in the right-click menu) and use the Label Text area to type the label, including line returns. The label can consist of multiple lines of text. You can also outline a label by choosing 'Rectangle' or 'RoundedRectangle' from the 'Outline' field in the properties panel
| Outlined label |
|---|
|
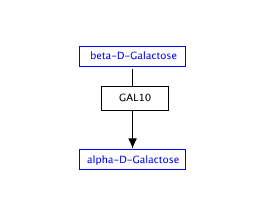
Representing metabolites
Metabolites and small molecules should not be represented as a label, but rather as a data object connected to a database entry (like genes and proteins). Use the 'Met' button in the editor to create a metabolite object.Double-click the object to fill in the textlabel, identifier and database, or simply search by name or identifier.
| Wrong | Right! |
|---|---|
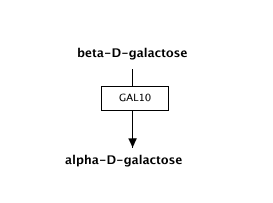
| 
|
Making Connections
Complicated connections involving curves or corners can easily be managed using "smart connectors". Any line or arrow can be made to curve or bend by choosing 'Curved' or 'Elbow' for 'Line type' in the right-click menu. Connect the ends of the line to their targets and the connector will bend where necessary. You can select the line again to adjust the waypoints of the path (blue diamond handles).
| Using elbows |
|---|

|
Replace arc/arrowhead combinations with curved connectors
| Using curves |
|---|

|
These options should help you style your pathway so that it is easy to "read" and has a layout that is familiar to any biologist or biochemist.
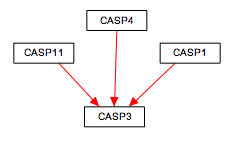
To make sure you've connected all your lines, press CTRL-L to reveal all unlinked line ends and highlight them in green.
| Before | After |
|---|---|
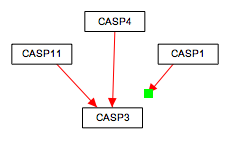
| 
|
Enforceable Policies
See policies page for list of rules regarding conduct, content and contributions at WikiPathways.